CTNet: A Convolutional Transformer Network for EEG-Based Motor Imagery Classification [Paper]
core idea: CNN (an improved version of EEGNet) + Transformer encoder
Our research builds upon and improves the EEG Conformer and EEG-ATCNet, and we sincerely thank the creators of these open-source project.
🎉🎉🎉 We've joined in braindecode toolbox. Use here for detailed info. Thanks to Bru and colleagues for helping with the modifications.
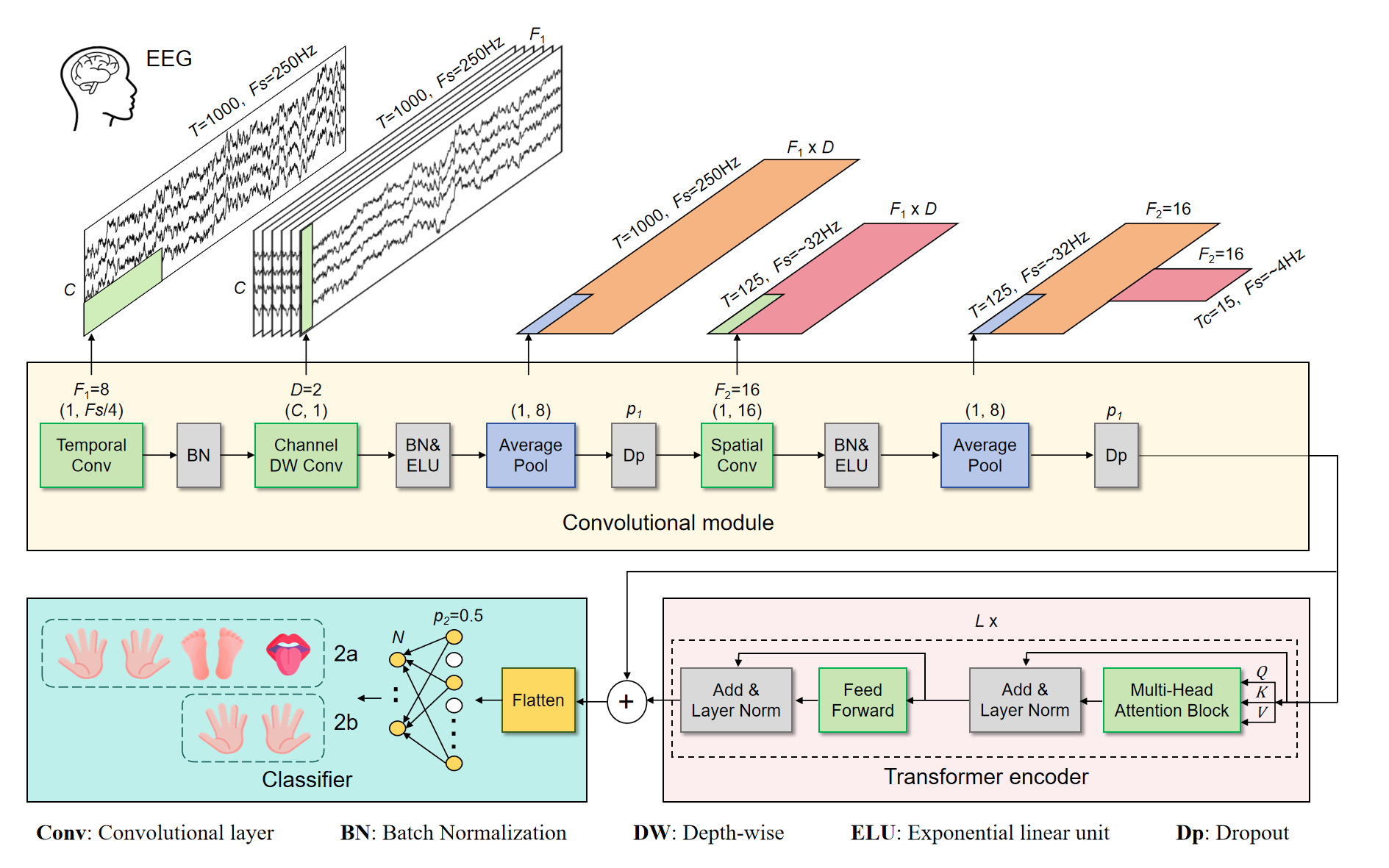
Brain-computer interface (BCI) technology bridges the direct communication between the brain and machines, unlocking new possibilities for human interaction and rehabilitation. EEG-based motor imagery (MI) plays a pivotal role in BCI, enabling the translation of thought into actionable commands for interactive and assistive technologies. However, the constrained decoding performance of brain signals poses a limitation to the broader application and development of BCI systems. In this study, we introduce a convolutional Transformer network (CTNet) designed for EEG-based MI classification. Firstly, CTNet employs a convolutional module analogous to EEGNet, dedicated to extracting local and spatial features from EEG time series. Subsequently, it incorporates a Transformer encoder module, leveraging a multi-head attention mechanism to discern the global dependencies of EEG's high-level features. Finally, a straightforward classifier module comprising fully connected layers is followed to categorize EEG signals. In subject-specific evaluations, CTNet achieved remarkable decoding accuracies of 82.52% and 88.49% on the BCI IV-2a and IV-2b datasets, respectively. Furthermore, in the challenging cross-subject assessments, CTNet achieved recognition accuracies of 58.64% on the BCI IV-2a dataset and 76.27% on the BCI IV-2b dataset. In both subject-specific and cross-subject evaluations, CTNet holds a leading position when compared to some of the state-of-the-art methods. This underscores the exceptional efficacy of our approach and its potential to set a new benchmark in EEG decoding.
Python 3.10
Pytorch 1.13.1
mne 1.5.1
datasets: BCI Competition IV-2a & IV-2b datasets
Create the BCICIV_2a_gdf directory to store the downloaded BCI IV-2a dataset.
Create the BCICIV_2b_gdf directory to store the downloaded BCI IV-2b dataset.
Create the true_labels directory to store the label data of the downloaded BCI IV-2a and 2b datasets.
Preprocessing dataset: Merge data and labels to create a dataloader for training the model (EDF -> MAT).
Run python3 preprocessing_for_2a.py for 2a dataset
Run python3 preprocessing_for_2b.py for 2b dataset
run python3 main_subject_specific.py or CTNet_2a_82.91.ipynb
run main_cross_subject_LOSO.ipynb
Under the LOSO setting, if data augmentation is required, it is recommended to refer to the updated training framework code, in which the data augmentation pipeline has been optimized: specifically, S&R performs data augmentation on each subject in the training set one by one, and then concatenates the data from different subjects to construct the training set. Refer to the LOSO code of TCANet. https://github.com/snailpt/TCANet/blob/main/LOSO_TCANet.ipynb
The original training set was split into training and validation subsets with a ratio of 7:3. Data augmentation was applied to expand the training set to three times (N_AUG=3) its original size.
Note: We observed that increasing the data augmentation factor (N_Aug in our code) leads to improved classification accuracy, but also results in a corresponding increase in training time.
Comparison of Subject-specific classification accuracy (in %) and kappa on the BCI IV-2a dataset.
| Method | Average±Std. | Kappa |
|---|---|---|
| ShallowConvNet | 75.69±11.76 | 0.6759 |
| DeepConvNet | 77.78±14.42 | 0.7037 |
| EEGNet | 77.39±12.47 | 0.6986 |
| TSF-STAN | 83.0±11.4 | 0.7650 |
| Conformer | 77.66±13.35 | 0.7022 |
| MI-CAT | 76.81±13.80 | 0.6920 |
| CTNet (Proposed) | 82.52±9.61 | 0.7670 |
Comparison of Subject-specific classification accuracy (in %) and kappa on the BCI IV-2b dataset.
| Method | Average±Std. | Kappa |
|---|---|---|
| ShallowConvNet | 85.13±10.74 | 0.7026 |
| DeepConvNet | 85.21±9.56 | 0.7042 |
| EEGNet | 87.71±9.33 | 0.7542 |
| TSF-STAN | 88.0±9.6 | - |
| Conformer | 85.87±10.73 | 0.7174 |
| MI-CAT | 85.28±12.93 | 0.7060 |
| CTNet (Proposed) | 88.49±9.03 | 0.7697 |
Comparison of cross-subject classification accuracy (in %) and kappa on the BCI IV-2a dataset.
| Method | A01 | A02 | A03 | A04 | A05 | A06 | A07 | A08 | A09 | Average±Std. | Kappa |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ShallowConvNet] | 66.84 | 46.53 | 67.53 | 52.26 | 34.38 | 39.76 | 65.45 | 71.18 | 66.84 | 56.75±13.77 | 0.4234 |
| DeepConvNet | 68.58 | 47.40 | 78.99 | 52.26 | 50.87 | 41.84 | 69.44 | 71.70 | 60.24 | 60.15±12.71 | 0.4686 |
| EEGNet | 69.79 | 42.01 | 79.51 | 50.87 | 35.76 | 37.15 | 65.80 | 67.36 | 63.37 | 56.85±15.82 | 0.4246 |
| Conformer | 68.75 | 37.33 | 69.62 | 43.58 | 29.51 | 35.24 | 58.33 | 74.48 | 63.89 | 53.41±17.08 | 0.3789 |
| CTNet (Proposed) | 69.27 | 43.92 | 79.34 | 55.38 | 43.92 | 36.11 | 65.10 | 70.66 | 64.06 | 58.64±14.61 | 0.4486 |
Comparison of cross-subject classification accuracy (in %) and kappa on the BCI IV-2b dataset.
| Method | B01 | B02 | B03 | B04 | B05 | B06 | B07 | B08 | B09 | Average±Std. | Kappa |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ShallowConvNet | 74.03 | 63.53 | 59.72 | 82.84 | 82.43 | 80.97 | 74.86 | 72.37 | 77.78 | 74.28±8.13 | 0.4856 |
| DeepConvNet | 74.03 | 65.15 | 63.47 | 80.81 | 82.70 | 74.86 | 81.39 | 76.32 | 77.92 | 75.18±6.84 | 0.5037 |
| EEGNet | 74.44 | 69.26 | 62.36 | 80.41 | 83.24 | 75.56 | 79.86 | 73.55 | 77.50 | 75.13±6.35 | 0.5026 |
| Conformer | 71.39 | 62.35 | 65.28 | 82.97 | 80.41 | 69.31 | 75.00 | 76.32 | 78.61 | 73.52±6.96 | 0.4703 |
| CTNet (Proposed) | 76.25 | 71.03 | 66.39 | 81.76 | 83.11 | 77.22 | 79.17 | 73.56 | 77.92 | 76.27±5.26 | 0.5252 |
Hope this code can be useful. I would appreciate you citing us in your paper. 😊
Zhao, W., Jiang, X., Zhang, B. et al. CTNet: a convolutional transformer network for EEG-based motor imagery classification. Sci Rep 14, 20237 (2024). https://doi.org/10.1038/s41598-024-71118-7
QQ discussion group (Motor imagery and Seizure Detection): 837800443
Email: zhaowei701@163.com